|
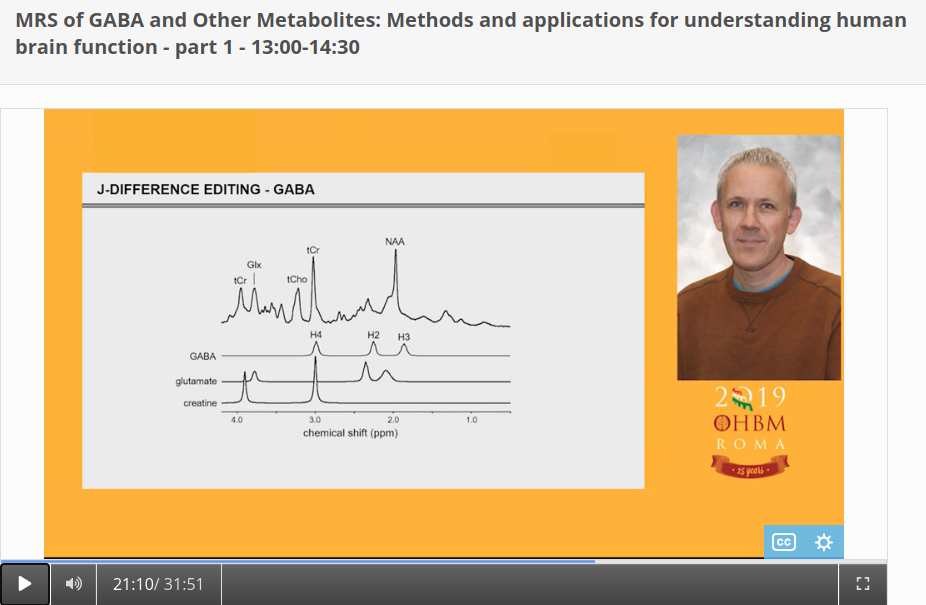
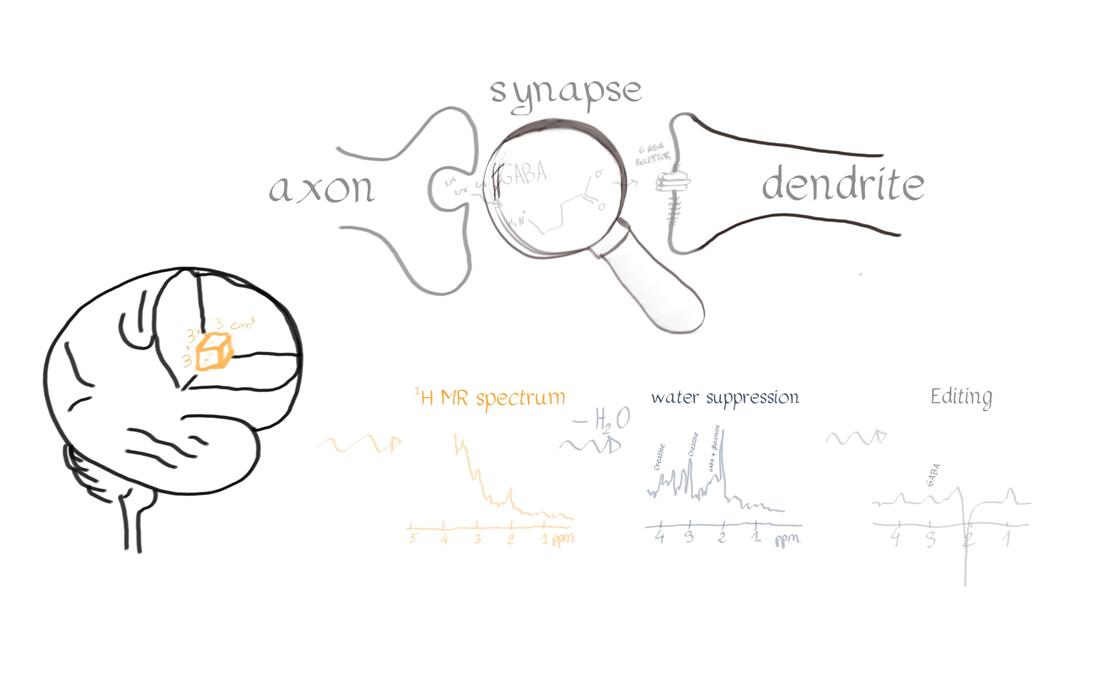
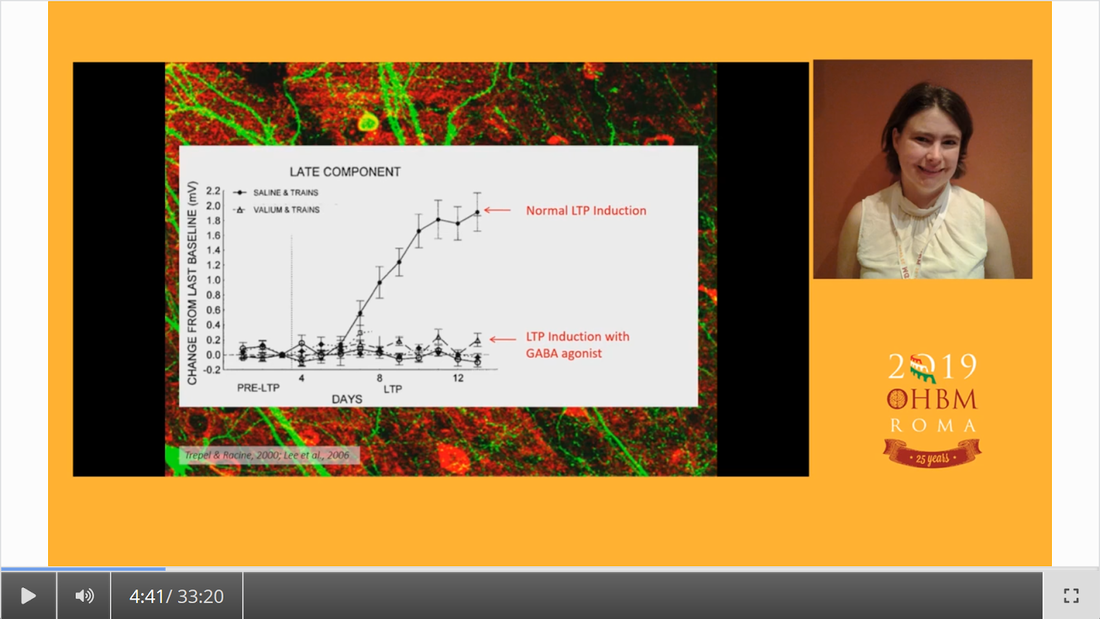
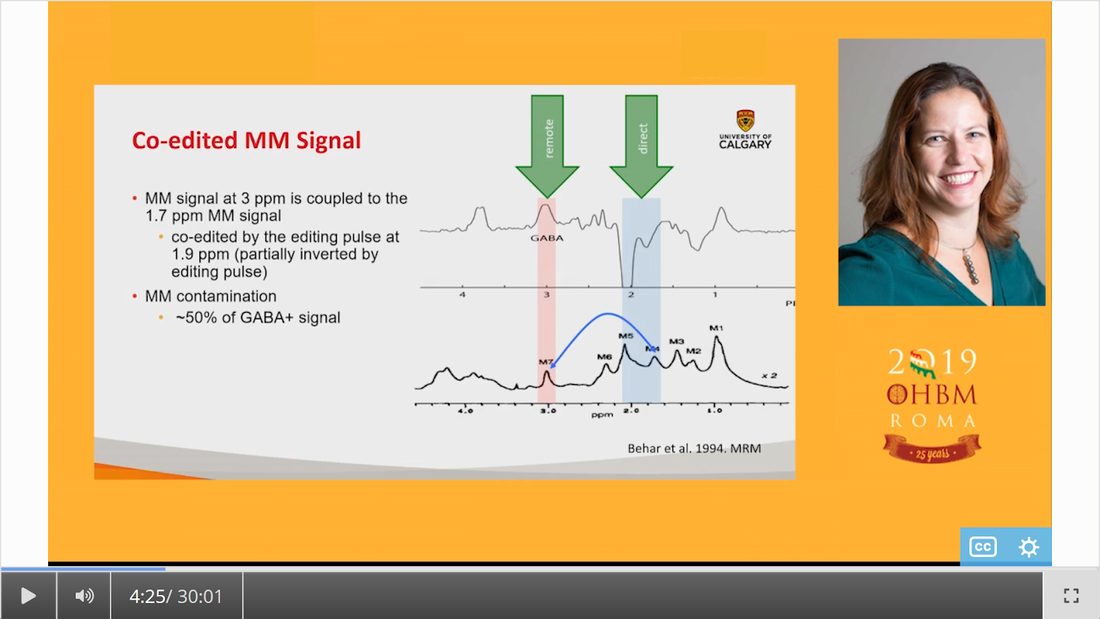
Authors: Katie Williams, Ilona Lipp, Mark Mikkelsen Infographic: Roselyne Chauvin Expert editors: James Kolasinski, Paul Mullins Newbie editors: Curtiss Chapman, Yana Dimech Introduction The noninvasive imaging tools that we Human Brain Mappers apply are most often being used to research brain structure and function. Neurotransmitter systems are something that we are aware of and use to take into account when coming up with hypotheses or interpreting our findings, but rarely make the direct subject of our investigation. Most of us have probably heard of GABA (gamma-aminobutyric acid) as the principal inhibitory neurotransmitter that is used by many interneurons. That we can also measure GABA in vivo with MR spectroscopy (MRS) is maybe less widely known. While this biomedical imaging tool opens many doors for neuroscience, measurement of GABA using MRS is not broadly used yet, possibly because special sequences and analysis methods are needed. At the OHBM Annual Meeting in 2019, for the first time, an educational session on GABA MRS was held. This post summarizes what was taught about the most important things you need to know if you’re considering GABA MRS for your research. Why should we care about GABA? As GABA is an inhibitory neurotransmitter, an intuitive way to think about it is that it can regulate neuronal firing, allowing the establishment of complex neural circuits and ensuring that the brain does not become “overactive”. This intuition is in line with the fact that drugs that act on the GABAergic system are traditionally used to treat anxiety and pain. In her video, Caroline Rae (from the beginning) emphasizes that when considering GABA, one should also consider glutamate, the excitatory neurotransmitter. She explains how GABA and glutamate are actively coupled at the synapse (min. 10:55). The ability of GABA to regulate glutamatergic firing makes it a neurotransmitter that is likely involved in many biological processes, one of them being brain plasticity, or the brain’s ability to structurally react to new situations (as explained by Charlotte Stagg from min. 1:55). Charlotte explaining the role of GABA in brain plasticity How can we measure metabolite concentrations with MRS? To understand the physical principles that give rise to MRS, it is helpful to take a few steps back to the basics. Robin de Graaf succinctly reviews (from min: 2:30) how nuclear magnetic resonance (NMR) in its essence is all about separation and detection of frequencies. In conventional MRI, we create a signal by interacting with the resonance frequencies of protons in a nucleus of interest – most typically that of hydrogen around 127 MHz at 3T and 298 MHz at 7T. MRS differs from typical MRI because it relies on something called the chemical shift effect. What does chemical shift mean? Depending on the chemical composition of a given molecule, the constituent protons experience different electronic shielding effects, resulting in slight differences in their resonance frequencies, which translates into many peaks appearing as an MR spectrum rather than a single clean signal peak at that free molecule’s resonance frequency. This is called chemical shift, because the biochemistry and environment of the molecule lead to a slight shift of its signal in the frequency domain. So, if one were to effectively “zoom in” on the hydrogen proton signal at 298 MHz, for example, we would see that the signal is actually composed of many smaller peaks in the range of a few hundred Hz surrounding this frequency in the MR spectrum. This signal reflects all MR-visible hydrogen-containing molecules in the sample. Since the water signal (coming from the hydrogen protons in the water molecule) is so much stronger in intensity than these other peaks, frequency-selective water suppression pulses are integrated into MRS pulse sequences to help reveal the less intense peaks that we are interested in. After showing us a zoomed-in shot of several peaks (screenshot below), Robin explains (from min. 4:00) how electronic shielding and chemical shift lead to consistent, exact locations of the peaks of different chemicals – or metabolites – in the MR spectrum. As resonance frequency depends on the field strength, Robin goes on to describe how moving away from frequency-based units to a parts-per-million (ppm) scale allows metabolite measurements to be more easily compared across field strengths (from min. 6:52). Robin showing us resonance frequency spectra of different nuclei Ok, what now? When combined with a pulse sequence with spatial localization, such as MEGA-PRESS or MEGA-(s)LASER (which Robin describes later, min. 17:56), a metabolite spectrum can be acquired from a volume of interest in the brain. The chemical shift effect holds true for all MR-detectable nuclei and, as such, for a long list of metabolites composed of those nuclei, including GABA. For this reason, many challenges that we face in measuring GABA concentrations apply universally in MRS. Clever use of relaxation properties and nuclear coupling effects give us a few solutions, however. Why is it challenging to measure GABA concentration with MRS? If specific metabolites like GABA can be measured with MRS, why are we not using it in every neuroimaging study? To be completely forthcoming, there are a number of challenges in conducting successful MRS measurements. Luckily, there are some options to deal with each of them. In spectroscopy, the signals we detect are very weak, so we have to run several hundred repeated acquisitions to obtain an acceptable averaged spectrum for quantification. Another way to boost SNR is to acquire spectra from larger voxels. Choosing an extra-large voxel size (by MRI standards) for higher SNR, however, is not an ideal solution because of heterogeneous tissue compositions in a voxel, and GABA concentration varies across different tissues. Ashley Harris explains that it is important to correct your measure for its tissue composition (from min. 8:53), because of known differences in GABA concentration in gray and white matter. Given the low SNR of metabolite signals, it has been common for a long time to use single-voxel MRS acquisitions. This is the reason that sometimes spectroscopy is not always categorized together with conventional MRI as a true brain imaging technique. However, using specialized pulse sequences, it is possible to acquire data from more than one region of interest using dual-voxel MRS, for example, which Muhammad Saleh describes in his video (min. 20:40). It is worth mentioning here that MRS imaging (MRSI, so spatially resolved MRS) approaches do exist, with which multiple voxels are acquired from a cubic volume, for example 3D MRSI can reach whole-brain coverage with a 14 × 14 × 12 voxel matrix size and 200 × 200 × 170 mm field of view (2.89 mL nominal voxel resolution), and technological advances to improve them are continually occurring. What is the problem with spectral overlap? What might be considered the biggest challenge for accurate metabolite measurements is spectral overlap. Given that so many biologically relevant molecules contain hydrogen protons, many with similar hydrogen structures, their signals will overlap, making it hard to get an accurate quantification of individual peaks that we care about, as Robin describes (from min. 9:30) in his video. If we cannot isolate the GABA peaks, then we cannot quantify them easily! In addition to the signals of identifiable metabolite peaks like creatine and glutamate that overlap with the GABA peaks in the spectrum, an underlying assortment of signals of broad peaks originating from macromolecules is present. (Here, macromolecules refer to a host of large molecules, including proteins, that differ from the smaller molecular structures such as GABA). The macromolecule (MM) signal is a biologically generic signal detected by in vivo MRS that usually consists of about ten peaks spread across the acquired spectrum. The MM signal can be attenuated using several acquisition solutions, which we describe below. However, it is important to note that the MM signals cannot be 100% removed, and their contribution is always present, to some extent, in a GABA measurement. There are several different options to approach spectral overlap, including moving to a higher field, like 7T, which improves the spectral resolution, meaning that the peaks are more spread out, and reduces the amount of overlap that occurs between them (an expensive solution, Robin notes, min. 10:50). Another possibility is to take advantage of T1 and T2 relaxation differences of different metabolites and use inversion recovery and spin-echo sequences in your experiments (as Robin describes, min. 11:46). What can we do about this spectral overlap problem? By far the most popular method for dealing with spectral overlap, and the most discussed technique for GABA quantification in the educational session, is the spectral editing approach: The same physical principles of nuclear interactions that make tiny changes to local magnetic environments and allow us to accomplish chemical shift imaging (i.e., to obtain spectra) offer a solution to spectral overlap. Nuclei that are chemically bonded to the same molecule, and thus generate multiple peaks for that molecule, are scalar-coupled, which, in quantum mechanics terms, means manipulation of one signal of a molecule also modulates the other signals of the same molecule. This phenomenon can be used to selectively manipulate overlapping signals and acquire the signal of interest. The figure above shows that GABA is composed of three major signals that are scalar-coupled to each other, and that glutamate and creatine have peaks overlapping in some locations. From min. 13:11 in his talk, Robin explains scalar coupling and how frequency-selective inversion pulses can be used during acquisition to modulate the signal of scalar-coupled molecules, but not the uncoupled ones. This is known as “editing” an MR spectrum. Using this technique, one can perform paired experiments, one with and one without the frequency-selective editing pulses, to recover the signal of the metabolite of interest. This technique, known as J-difference editing, is a powerful MRS technique used for measuring GABA in the brain. It should be noted that while scalar coupling helps us to more specifically acquire our signal of interest, co-editing always occurs, and attention should be paid about which molecules are being inverted. Robin describes a simple pulse sequence for a full J-difference editing experiment, using GABA as an example (from min. 17:56), while Muhammad Saleh speaks extensively in his video about special GABA editing sequences and ways to speed up editing experiments to increase the information extracted from the data acquired. And that brings us back to the topic of challenges in measuring GABA: applying solutions to acquire good spectra significantly increases scan duration, giving rise to more temporal instabilities in the signal, specifically frequency offsets. Frequency offsets are shifts in the main magnetic field that most often occur either because of heating/cooling of the gradient hardware elements in the scanner or bulk participant head motion. In her talk, around 24 minutes into the video, Ashley discusses this problem and how sometimes it can be fixed retrospectively through frequency alignment. Robin explaining the J-difference strategy of measuring GABA What do I need to consider when setting up a GABA-edited MRS acquisition? There are some essential questions to answer when setting up a GABA-edited MRS experiment. Of course, the first is where in the brain you want to measure GABA. For hypothesis-driven studies, this will be determined either by the functional neuroanatomy of the aspect of brain function being studied or by the regions implicated in the neuropathology/pathogenesis of a particular brain disorder/disease. It is worth reiterating that, given the low SNR of the GABA signal, the size of the volume of interest will be on the order of cubic centimeters. Thus, one will need to be aware of the limitations on the specificity of where in the brain GABA will be measured. For GABA editing, voxels tend to have around 27 mL tissue in them (e.g., 3 × 3 × 3 cm3) in volume to attain reasonable SNR. In the MRS literature, the voxel size is often reported in volume, as this is the relevant factor for SNR. Oh, such large voxels!? Can I not just go to 7T and get a better spatial resolution? Should it be possible (and desired) to perform MRS experiments at ultra-high field (>3T), then the benefits of a higher field strength would alleviate some of the challenges of MRS acquisitions. Aside from the associated increase in inherent SNR of metabolite signals, and the already mentioned increase in spectral resolution (the separation of peaks in the spectrum), high field measurements allow improved selectivity of editing pulses. These advantages of ultra-high field MRS make it more feasible to detect GABA without using editing. Nevertheless, editing at 3T remains the most commonly used approach for measuring GABA that you will encounter in the literature. Another consideration for increasing SNR is scan duration. In edited MRS, each acquisition is repeated (usually several hundred times) in order to perform signal averaging to improve the SNR of the detected metabolite signals. As Ashley Harris explains in her presentation (from min. 19:43), the question of how many averages are needed (how many times to repeat the measurement in one scan acquisition),(i.e., how long to scan) will depend on voxel size, the scientific question being asked, and the region in which you are scanning. Some regions like the occipital lobe provide good SNR and therefore allow you to scan for shorter periods. In contrast, other regions like the temporal lobe are more challenging to acquire high-quality data in and necessitate collecting relatively more averages. Anything else I need to think about? It is also worth considering the order in which you run your different MRI/S acquisitions in a given scan protocol. When conducting a study, it is quite likely you will be acquiring a variety of scans, such as fMRI, diffusion MRI, and MRS. Sequences that involve rapid switching of gradients (e.g., EPI and DWI) will lead to heating and subsequent cooling of the scanner’s hardware elements. This causes shifts (or drift) in the B0 field (and thus its frequency) that can have a considerably detrimental effect on edited MRS acquisitions, which require frequency stability to ensure the narrowband frequency-selective editing pulses perform as intended. Performing MRS acquisitions before any scans that make use of high gradient duty cycles can help lessen the impact of frequency drift on acquisition performance. Also, the use of prospective and retrospective frequency alignment methods can mitigate the detrimental effects of frequency drift on spectra. Ashley also talks about this in her presentation (from min. 22:30). How do I know whether the quality of my spectra is good enough? Several signal artifacts can lead to poor quality of MRS data. An excellent place to start is by reading this paper, which describes in detail the kind of artifacts one would see in corrupted MRS data. A full description of artifacts is beyond the scope of this blog post. Still, one thing in particular that can significantly degrade the quality of your spectra is participant motion. The comparatively longer scan times of edited MRS acquisitions, unfortunately, provide more opportunities for a participant to move and worsen spectral quality. Some simple steps that can be taken to prevent motion artifacts include emphasizing to participants the importance of remaining as still as they reasonably can when they hear the scans running and acquiring structural/fast localizer images and MRS data consecutively so that voxels are placed as accurately as possible given participants’ current head position. The act (art) of rating the quality of MR spectra can be challenging to those new to MRS. Since MRS is methodologically distinct in several important ways from MRI, quality analysis may be less intuitive to new users who are more familiar with the latter technique. Typically, a good approach to quality analysis (when possible) is to consult a colleague (internally or externally) who has experience with MRS. A review of 2–3 pilot datasets can go a long way to establishing the predicted quality of MRS data for a proposed study. When an investigation is underway, it is highly beneficial to review data as they are collected. Continual reviews of data can prevent situations where a series of datasets have been acquired with significant artifacts that would lead to their removal from further analysis, which could potentially seriously undermine the success of a study. Ok, so now I have a spectrum, but how do I quantify GABA? Once you have acquired some GABA-edited MRS data, you can quantify the GABA from the data you have collected. There are some software analysis packages available to users that can quantify GABA from edited MRS data. These include Gannet, jMRUI, LCModel, TARQUIN and, most recently, also FSL. Each has its own strengths and weaknesses and particular learning curve, but each will allow you to derive a quantified measurement of GABA from your MRS data. The GABA signal is either quantified in the time or (more commonly) the frequency domain, where either the amplitude or the area of the GABA signal is used to determine the concentration (as concentration is proportional to signal amplitude or area). While a description of each package is beyond the scope of this blog, readers are advised to read the following papers for further information: Edden RAE, Puts NAJ, Harris AD, Barker PB, Evans CJ. Gannet: A batch-processing tool for the quantitative analysis of gamma-aminobutyric acid-edited MR spectroscopy spectra. J Magn Reson Imaging. 2014;40(6):1445-1452 Provencher SW. Estimation of metabolite concentrations from localized in vivo proton NMR spectra. Magn Reson Med. 1993;30(6):672-679 Stefan D, Cesare F Di, Andrasescu A, et al. Quantitation of magnetic resonance spectroscopy signals: the jMRUI software package. Meas Sci Technol. 2009;20(10):104035 Wilson M, Reynolds G, Kauppinen RA, Arvanitis TN, Peet AC. A constrained least-squares approach to the automated quantitation of in vivo 1H magnetic resonance spectroscopy data. Magn Reson Med. 2011;65(1):1-12 Ok, so now I have a GABA concentration measure. But what does this measure mean? There are two things to consider. First is the specificity of the GABA signal we are detecting. As Ashley describes (from min. 3:25), the GABA signal is contaminated by a co-edited MM signal that leads to the observed GABA signal at 3 ppm being ~50% MM. For this reason, it is conventional in the field to define edited GABA measurements acquired by standard editing as “GABA+” measurements, to acknowledge the MM contribution. The majority of studies in the literature applying GABA-edited MRS will have acquired measurements of this sort. It is, therefore, important to recognize this limitation when setting up an experiment (it may impact the interpretation of your findings). Alternatively, one may implement an MM suppression technique that removes the MM signal underlying the GABA+ signal so that measurements are a “purer” measure of GABA. However, this comes at the cost of reduced SNR (from min. 5:00). Ashley warning us that we are not only measuring GABA The second thing to consider is that MRS measurements of GABA are not direct measurements of neuronal inhibition. In her video, Caroline describes that there are actually four types of inhibition (from min. 8:43), several metabolic pathways for GABA, and multiple GABA receptor types (from min. 11:49). Based on the intuitive way of thinking about GABA as a inhibitory neurotransmitter, one may expect to find negative relationships between GABA levels and brain activity, such as measured with fMRI (also see this paper for guidance for how to form and test hypotheses about the relationship between neurochemistry and activity). However, when it comes to energy expenditure and metabolism that underlie functional imaging measures, such as BOLD signal changes, things are not so simple (see this paper to show that a relationship between GABA and BOLD is not easy to find). Caroline explains how excitatory and inhibitory activity together can either increase or decrease energy metabolism, depending on the context (min. 16:36), and even more, GABA can directly modulate blood flow (from min. 22:50). Therefore, the interpretation of GABA levels, measured with MRS, is far from straightforward. Caroline (from min. 23:20) points out that the measures reflect neurotransmitter and metabolic pools, they are dependent on brain energy and activity, and they could reflect tonic inhibition. As head motion, different types of medication, and tissue composition of your voxel all can have an impact on the outcome measure, Nicolaas (from min. 27:03) recommends considering these confounding factors in your analysis and data interpretation. Additionally, menstrual cycle and time of the day have been found to be potential influencers of MRS-measured GABA concentration.
Does the uncertainty of interpretation not mean that it is pointless to do GABA MRS? We understand that the difficulty in interpretation may be off-putting. But at this point, we want to remind you that most of the imaging measures we look at are indirect. Think about BOLD as a measure of neural activity for example. These indirect measures are still useful for inferring something about a clinical condition, and in combination with other methods, develop a more holistic picture of what is going on. In his video, Nicolaas gives a number of examples for how GABA MRS has been used in clinical research, such as in neurodevelopmental disorders (from min. 7:46), depression (min. 9:44), personality disorders (min. 11:34) and schizophrenia (min. 13:40). Due to its role in learning and plasticity, GABA MRS has also been used in healthy populations. In her video, Charlotte Stagg provides some examples of how GABA, measured by MRS, changes in perceptual learning (from min. 10:00), overlearning (from min. 16:43), and learning how to juggle, as an example of long-term learning (from min. 20:06). Ok, so if I do want to start using GABA MRS in my research, how can I learn more? A good place to start is to read these overview/consensus papers: Bogner W, Hangel G, Esmaeili M, Andronesi OC. 1D-spectral editing and 2D multispectral in vivo 1H-MRS and 1H-MRSI - Methods and applications. Anal Biochem. 2017;529:48-64 Harris AD, Saleh MG, Edden RAE. Edited 1H magnetic resonance spectroscopy in vivo: Methods and metabolites. Magn Reson Med. 2017;77(4):1377-1389 Mullins PG, McGonigle DJ, O’Gorman RL, et al. Current practice in the use of MEGA-PRESS spectroscopy for the detection of GABA. Neuroimage. 2014;86:43-52 Another excellent resource is Robin’s book (which can be downloaded if your institution has access): In Vivo NMR Spectroscopy: Principles and Techniques Finally, the MRS community has recently begun assembling a curated collection of resources for data acquisition and analysis in the form of MRSHub. The forum is a great place to pose questions that can be answered by experts.
0 Comments
Your comment will be posted after it is approved.
Leave a Reply. |
BLOG HOME
Archives
January 2024
|


 RSS Feed
RSS Feed