Lavinia Carmen Uscatescu and Sin KimThe third entry in our 2023 Keynote Presenter series  Dr. Emma Robinson is a Senior Lecturer (Assoc. Professor) at King’s College London. Her development of the Multimodal Surface Matching (MSM) software for cortical surface registration has been instrumental to the development of the Human Connectome Project’s multimodal parcellation of the human cortex. She is currently developing interpretable machine learning models for personalized prediction of disease progression. In this interview, Dr. Robinson describes the advantages of interpretable machine learning models and the methodological challenges she faced during the development of this framework. Her approach to identifying disease-related changes in individual brain scans attempts to circumvent two limitations of traditional approaches: (1) the over-reliance on population averages; and (2) the opacity of “black-box” machine learning algorithms such as deep neural networks. However, her extensive experience working on the Human Connectome Project led her to realize that traditional image registration methods may not be sufficient for individualized predictions. In addition, Dr. Robinson shared how her relationship with her mentors shaped the trajectory of her current career. Her mentors not only guided her on the application of computational methods to neuroscience but also encouraged her to develop her own methods. At OHBM 2023, Dr. Robinson will present how her work contributes to improved personalized predictions of cortical features in patient populations and how interpretable machine learning approaches can enhance precision. You can find the video interview here and listen to the audio-only podcast version here (or on your podcast app of choice).
0 Comments
Elisa Guma and Simon SteinkampContinuing our OHBM2023 keynote interview series  Dr. Emily Jacobs is an Associate Professor of Psychological & Brain Sciences and the director of the Ann S. Bowers Women’s Health Initiative at University of California, Santa Barbara. She received her PhD in Neuroscience at the University of California, Berkeley, and her BA in Neuroscience from Smith College. Prior to UCSB, she was an instructor at Harvard Medical School and at the Department of Medicine/Division of Women’s Health at Brigham & Women’s Hospital. In this interview, we discuss the pioneering work of Dr. Jacobs and her group in leveraging brain imaging, computation, and endocrine approaches to deepen our understanding of the influence of sex hormones on the central nervous system across spatial and temporal scales. Her group uses structural and functional neuroimaging methods to explore how the brain changes in response to endogenous hormonal changes, such as across the menstrual cycle, pregnancy, or menopause, as well as to exogenous hormones via oral hormonal contraceptives. Through the Ann S. Bowers Women’s Health Initiative, Dr. Jacobs and her group are working towards creating a population-level brain imaging dataset to advance our understanding of women’s brain health across the lifespan. Dr. Jacobs also shares her journey into neuroscience research, her thoughts on how science can inform public policy, and talks about her groups’ efforts to improve girls’ representation in STEM by partnering with K-12 groups. This work was featured in the book STEMinists: The Lifework of 12 Women Scientists and Engineers. At OHBM 2023, Dr. Jacobs will highlight the power of sex steroid hormones and the role that they play in shaping the brain over multiple timescales, drawing attention to some of the reasons why it has taken the field so long to focus on women’s brain health. You can find the video interview here and listen to the audio-only podcast version here (or on your podcast app of choice). What to Expect from the Diversity and Inclusivity Committee at The 2023 OHBM Annual Meeting6/13/2023 Alexander Barnett, Christienne Damatac, Eduardo A. Garza-Villarreal, Julia Kam, Lucina Uddin, and Maryam ZiaeiOn behalf of the OHBM Diversity & Inclusivity Committee At this year’s OHBM meeting in Montreal, cutting-edge technology and a commitment to diversity and inclusivity will converge in an array of events curated by the Diversity and Inclusivity Committee (DIC). We will showcase the transformative power of thoughtful methodology in enhancing accessibility to and fostering a sense of belonging in neuroscience and neuroimaging.
First, the 5th annual DIC symposium will feature a panel of experts on groundbreaking technological solutions for supporting our diverse global community. From revolutionizing accessibility for individuals with visual and auditory impairments to promoting inclusivity in neuroimaging studies, this symposium promises to inspire us to actively improve our own research to be more inclusive. Next, at the Multilingual Kids Review, we will engage young reviewers from diverse backgrounds in critically assessing a scientific presentation. Finally, the Diversity & Inclusivity Roundtable will focus on advancing diversity across multiple dimensions, examining strategies for organizing diverse symposia, educational courses, and brain hackathons. We hope you will join us as we explore how technology, diversity, and inclusivity intersect to shape the future of neuroscience and neuroimaging. Together, we can work towards creating a welcoming environment for OHBM’s diverse and international membership. Xinhui Li and Kevin SitekKicking off the 2023 Keynote Lecture Series
By Charlotte Rae, Nikhil Bhagwat, Peer Herholz, Irene Faiman, and Niall DuncanFROM THE SUSTAINABILITY AND ENVIRONMENT ACTION SIG (SEA-SIG) As we prepare for the 2023 annual meeting in Montreal, many of us have started looking into travel arrangements, accommodation, and generally getting ready for the annual meeting. From booking travel to planning your time in Montreal, there are lots of ways that you can make your 2023 meeting experience more sustainable. Here we highlight some often overlooked tips, from getting around the city to poster printing and more
Yohan YeeA look back before moving forward to 2023 keynote interviews Prof Janaina [Jana-eena] Mourao-Miranda leads the Machine Learning and Neuroimaging Lab within the Centre for Medical Image Computing (CMIC) at the Department of Computer Science, University College London (UCL), where she applies pattern recognition and machine learning techniques to neuroimaging data. A major theme within Prof Mourao-Miranda’s research is uncovering the relationship between brain and behaviour.
At OHBM 2022, Prof Mourao-Miranda gave a keynote lecture on machine learning in neuroimaging and psychiatry. You can find a recording of Prof Mourao-Miranda’s talk here. Below is an edited transcript of an interview conducted with Prof Mourao-Miranda on June 17, 2022. Jean ChenOn behalf of the Women in OHBM Special Interest Group Over half of OHBM conference attendees are women scientists (according to data from OHBM Executive Staff), and many of them continue to face gender-specific barriers in their work as human brain mappers. To enable members to learn from one another and mutually support career success, we created the Women in OHBM Special Interest Group (SIG), officially recognized as of March 2023.
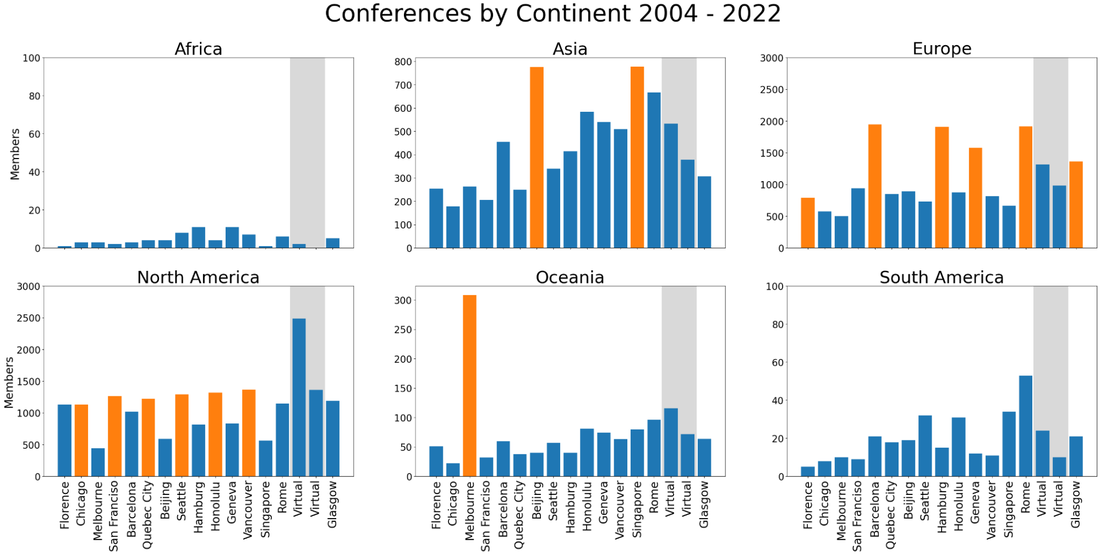
The product of a 2-year informal consultation process, the Women in OHBM SIG was motivated by a group of mid- and early-career OHBM women scientists with shared interests and challenges around gender equity in terms of scientific impact, career development and work-life balance, among other topics. The SIG aims to provide a community for OHBM women to network, promote mutual career development and facilitate scientific exchange amongst women scientists. Simon SteinkampLooking at OHBM membership data and a new membership tier OHBM transitioned to a scientific society in 2018 with the goal of delivering year-round engagement for the society’s members rather than being solely about the annual meeting. However, if one looks at the raw membership numbers over the last 17 years, the main perceived membership benefit remains connected to the annual meeting. In particular, yearly membership numbers fluctuate with meeting location, with highest membership numbers from the same geographic region. Unfortunately, this means that the society-based aspects of OHBM and many of the benefits of being an OHBM member have been largely overlooked. Thus, OHBM leadership is introducing a new MEMBERSHIP+ initiative to create a greater engagement of members on top of the annual meeting, strengthening OHBM as a society. Alfie WearnOn behalf of the OHBM Communications Committee Podcast Team
Naomi L. Gaggi
Beth Slater, with support from the OHBM Executive Office Happy New Year and welcome to 2023, the year that OHBM will travel to Montréal for the 29th Annual Meeting. This meeting will be primarily held in person—but in a change from previous years, all members of OHBM can upload approved content and automatically have access to Annual Meeting content in a virtual space, regardless of registration status. In previous years, presenters were required to register for the Annual Meeting prior to uploading their content. New this year, presenters who are unable to participate in person may upload content simply by being a member of OHBM. This shift will reduce the financial burden for individuals who cannot travel to Montréal. Membership can be renewed here at any time.
Xinhui Li, Lena Oestreich, Aman Badhwar, Sridar Narayanan, Gladys Heng, Robin GutzenOn behalf of the OHBM Brain-Art Special Interest Group
OHBM Blog TeamHappy holidays! Well, it’s here—the end of 2022. With OHBM’s first in-person annual meeting since before the COVID-19 pandemic, 2022 shaped up to be a busy year for the OHBM Communications Committee! We gathered up the experiences and thoughts of this year’s OHBM blog contributors to hear how everyone’s doing and what they’re looking forward to in 2023. (For a blast from the past, here are the 2021, 2020, and 2019 posts.)  Kevin Sitek, Blog team lead and Committee chair-elect https://sitek.github.io/ I am very thankful that 2022 brought the return of mostly normal activities for me, particularly international travel and an in-person OHBM annual meeting! It was amazing seeing so many old friends and collaborators (and meeting plenty of new ones) in Glasgow. I was also able to visit a few other cities before and after the conference, which scratched a two-plus-year travel itch. Within the Communications Committee, I shifted to more behind-the-scenes activities in 2022, but not before flipping the microphone on the podcast host and interviewing Peter Bandettini on the Neurosalience podcast. As blog team lead, I know there’s a ton of great content coming in the next year. Thanks for reading and listening along with us, and we hope to see you in 2023! Alfie Wearn
OHBM Diversity and Inclusivity CommitteeEvery year the OHBM Program Committee takes on the challenging task of creating content for the annual meeting that appeals to the multifaceted, global OHBM community. One of the top priorities for the committee is to ensure diversity of presenters at the meeting. However, it may be unclear how to achieve this goal.
Currently, the submission guidelines for symposia and educational courses state that submissions should provide a “statement on presenter diversity.” We hope to provide a discussion of what the statement of presenter diversity means and how organizers can ensure that a symposium submission meets this requirement. The Diversity and Inclusivity Committee has some ideas that we hope will move this discussion forward and provide concrete guidelines. Alfie Wearn(new) Podcast team lead
Kevin SitekBlog team lead and ComCom chair-elect
With such a dramatic change to the upcoming calendar, it’s critical for the human brain mapping community to know what to expect for OHBM 2023 so that they can plan for the new schedule. To that end, we communicated with Alex Shun (Communications Manager at the OHBM Executive Office), Michele Veldsman (OHBM Council Secretary), and Michel Thiebaut de Schotten (OHBM Council Chair) to discuss the reasons for the change, the decision-making process, and the issues and opportunities that arise from this date shift.
Why have the dates of OHBM 2023 in Montreal changed, and why was the decision made at this point? Michel Thiebaut de Schotten (MTdS): The Canadian Grand Prix is typically held in June in Montreal, but until very recently we didn’t know dates—it’s usually earlier in the month. When the overlapping event dates were announced last week, we knew it would be a big problem for accommodations (since around 300,000 people visit Montreal for the Grand Prix). Michele Veldsman (MV): The Grand Prix overlap pushed the prices of everything up three-fold—which makes it completely inaccessible for most people. Alex Shun (AS): The overall attendee experience was the driving force behind the OHBM 2023 date change. The Executive Office was in close contact with our local vendors as we awaited the Grand Prix schedule to be publicized and worked to create a solution when we learned of the overlap. Accommodation prices, flights, social venue costs, and the overall ease of getting around the city would have hugely impacted our community and we wanted to ensure our attendees have a positive experience in Montreal. Alexander HolmesPhD Candidate at the Neural Systems and Behaviour Lab, Monash University, Australia  Dr. Juan (Helen) Zhou is an Associate Professor and Principal Investigator of the Multimodal Neuroimaging in Neuropsychiatric Disorders Laboratory in the Centre for Sleep and Cognition, Yong Loo Lin School of Medicine, National University of Singapore (NUS). She also holds a joint appointment with the Department of Electrical and Computer Engineering, NUS, and she currently serves as the Deputy Director for the Centre for Translational Magnetic Resonance Research at Yong Loo Lin School of Medicine. She recently finished her term as Council Secretary and a member of the Program Committee of the Organization for Human Brain Mapping. Across these roles, Dr Zhou’s research focuses on the network-based vulnerability hypothesis in disease. Specifically, her lab studies the neural bases of human cognitive functions and the associated vulnerability patterns in ageing and neuropsychiatric disorders using multimodal neuroimaging methods, psychophysical techniques, and machine learning approaches. Dr. Zhou presented a keynote address at OHBM 2022 in Glasgow—read on to learn about her research, career path, and hopes for the future of neuroimaging! Alexander Holmes (AH): Welcome Dr. Zhou, thank you so much for joining us here—it is an honour to have you with us. Can you first tell us about your pathway into science and how you got to where you are now? Helen Zhou (HZ): Ah, do you want the short answer or the long answer? When I was doing my undergraduate studies at the School of Computer Science and Engineering in Singapore, I was a part of this accelerated Masters program. During our final year, we needed to do some research, which was where I became interested in algorithms, neural networks, and image processing. When I tried these machine learning projects, it was my first hands-on experience using these algorithms to solve real problems. So, there were many ups and downs (Laughs). Anastasia BrovkinPhD Candidate in Computational Neuroscience, Universitätsklinikum Hamburg Eppendorf
Yohan Yee, on behalf of the Communications CommitteeAre you interested in sharing new research and ideas within (and beyond!) the human brain mapping community? Do you want to be more involved in OHBM and learn about the exciting research led by our community members?
Then apply to join the OHBM Communications Committee! We’re currently accepting applications for new team members through 15 August (5pm PDT). Read on to discover what the Communications Committee does and how you can get involved. Steven BaeteAssistant Professor of Radiology, Center for Biomedical Imaging, New York University Dr. Jonathan Polimeni is Assistant Professor of Radiology at Harvard Medical School and of Biomedical Engineering at Massachusetts General Hospital, Athinoula A. Martinos Center for Biomedical Imaging. In his research, he focuses on the fundamental understanding of neural activity in the brain, often in the visual cortex. In pursuing this understanding, Dr. Polimeni has along the way pushed the boundaries of fMRI. His work has resulted in many contributions to both neuroscience and functional imaging science, both in insights gained and in technical advancements. We had the opportunity to chat with Dr. Polimeni about his experience as a scientist and his vision on functional imaging. Steven Baete (SB): To start things off, if you were not talking to brain mappers or scientists, how would you describe your research and your most proud scientific accomplishment?
Jon Polimeni (JP): I would first say that MRI tracks brain function not by detecting neural activity directly. Instead, you can see where the blood flow is increased in the brain in order to deliver oxygen to where it is needed. And because of the magnetic properties of the blood, we can track this with MRI. The blood vessels of the brain are quite smart, and can deliver blood exactly to where it is needed, when it is needed. The goal of my work is to understand how the blood flow is delivered to the brain and to build technologies to image this delivery more clearly. To make functional MRI a better tool to see neural activity and brain function in working brains. My proudest scientific accomplishment is just to be able to contribute. As a domain, I feel like we have been able to both develop technologies to improve our abilities to track brain function with fMRI and to shed a few insights into this blood flow regulation. I am not sure if I can point to a single achievement, I am just happy to be a part of this endeavor. Rahul Gaurav Movement, Investigations and Therapeutics (MOV'IT) team and the Center for NeuroImaging Research (CENIR) at the Paris Brain Institute (ICM - Institut du Cerveau), Sorbonne Université, INSERM U1127, CNRS UMR 7225, Pitié-Salpêtrière Hospital, France. Dr Lozano is a neurosurgeon and University Professor at the University of Toronto, where he is best known for his work in the field of Deep Brain Stimulation (DBS) and Magnetic Resonance-guided Focused Ultrasound (MRgFUS). His team has mapped cortical and subcortical circuits in the human brain and has advanced novel treatments for Parkinson’s disease and for depression, dystonia, anorexia, Huntington’s, and Alzheimer’s disease. Dr. Lozano has over 750 publications and serves on the boards of several international organizations. He has trained over 70 international postdoctoral fellows. He has received a number of honors including Doctor Honoris Causa from the University of Sevilla, the Olivecrona Medal, the Pioneer in Medicine Award, and the Dandy Medal. He has been elected to the Royal Society of Canada, has received the Order of Spain, and is an Officer of the Order of Canada. Here, he sits down to discuss his work and his OHBM2022 Talairach address.
OHBM2022 Keynote Interview with Sarah Genon: Mapping the steps in the brain-behavior tango6/12/2022 Gopi Deshpande Professor of Electrical and Computer Engineering at Auburn University “I was a lifelong chain smoker. Nothing in the world could stop me from smoking. Then, one day I had brain injury and my Insula was damaged. When I woke up, it felt like the urge to smoke had suddenly disappeared. It was as if a switch had been turned off. I could not believe what I was experiencing” - By an anonymous ex chain smoker
Fakhereh Attar PhD student at the Max Planck Institute for Human Cognitive and Brain Sciences
Diversity & Inclusivity Events at the 2022 OHBM Annual Meeting: If you want to go far, go together5/27/2022 Valentina Borghesani, Lucina Uddin, Kangjoo Lee, Aman Badwhar & Rosanna OlsenOn behalf of the OHBM Diversity & Inclusivity Committee The last two years have brought new challenges for the members of our global OHBM community. In particular, the COVID-19 pandemic highlighted and exacerbated existing inequities in having access to healthcare, including vaccines. During this time, we have also witnessed persistent racial and/or ethnocultural discrimination, which continues to affect members of our Society around the globe. Finally, while our recent survey indicated that many of our members have reported positive changes that support Diversity and Inclusivity at OHBM in the past two years, there is still a major lack of geographical representation within our Council and at our annual meeting. We, the OHBM Diversity & Inclusivity Committee, will continue to shine a light on these issues by discussing existing barriers and proposing solutions at the OHBM meeting Diversity Symposium and Roundtable events. We also continue to engage the “scientists of the future” around the world in our 2nd annual multilingual Kids Live Review (virtual) events.
|
BLOG HOME
Archives
January 2024
|














 RSS Feed
RSS Feed